Scientific Research
My scientific expertise lies in the fundamental biology of CRISPR-Cas immune systems, CRISPR-based genome-editing technologies, and applications of genetic engineering, especially in medicine.
Once a scientist, always a scientist
In my current role at Arcadia Science, we don’t publish in academic journals—we are pushing ourselves to reimagine what open, effective scientific publishing should look like.
Scientific Training
-
Arcadia aims to develop therapeutics inspired by evolution. We search agnostically across the tree of life to find hidden biological innovations that can improve human health.
I’m leading Arcadia’s open science approach and making research communications more expedient, transparent, and useful.
-
At the IGI, I focused on the world of genome editing. I stopped doing experiments, but tracked all the latest developments in CRISPR research, genetic engineering, applications, and societal issues.
I was lucky to work with experts across a range of disciplines within life, physical, and social sciences, greatly expanding my perspective.
-
I got my PhD from UC Berkeley, working in Jennifer Doudna’s lab. I performed biochemical and structural studies of CRISPR-mediated bacterial immunity. My focus was on I-C and I-E multi-subunit effector complexes.
I also had a front row seat for the “CRISPR revolution” and got a crash course in genetic engineering.
-
I studied RNAi in a parasite, oncogenic viral proteins, and yeast prions in the labs of Elisabetta Ullu, Dan DiMaio, and Tricia Serio, respectively.
These wide-ranging experiences sparked my love of research and revealed to me that there are a variety of ways of thinking about science. I’m forever grateful to my early mentors!

Publications

Reimagining Scientific Publishing
At Arcadia Science, we share our research directly with the scientific community on a platform called PubPub, without journals or other forms of gatekeeping. The entire effort is an experiment in and of itself. I work with all of our scientists to communicate their science, and more broadly, to design and test this new publishing model. We’re tracking our goals and progress in an overall project narrative and providing in-depth updates in individual pubs, listed below.
The experiment begins: Arcadia publishing 1.0 [Perspective]
Building on the open-source platform PubPub, we’re sharing the first iteration of our publishing website. In addition to posting our first set of research pubs, we’re documenting our progress in developing this new system for sharing science and hope you’ll provide feedback.
Avasthi P & Hochstrasser ML. Arcadia Science (2022)
The experiment continues: Arcadia publishing 2.0 [Perspective]
Two years into our publishing experiment, we’ve learned a lot. We built internal processes that worked but inadvertently decreased scientists' agency and creativity. Now, we’re minimizing process in an effort to empower our scientists to share their work how they see fit.
Avasthi P, Hochstrasser ML, Roth R. Arcadia Science (2024)
Early update on Arcadia publishing 2.0: Scientists are in charge, speed is an issue [Result]
Since starting v2 of our publishing model to restore scientist agency, pub “quality” has been similar, and this approach is more efficient overall. The major downside is that pubs take longer. We’re pursuing solutions to this and other problems but feel we’re on the right track.
Avasthi P, Hochstrasser ML, Roth R. Arcadia Science (2024)
CRISPR Biology
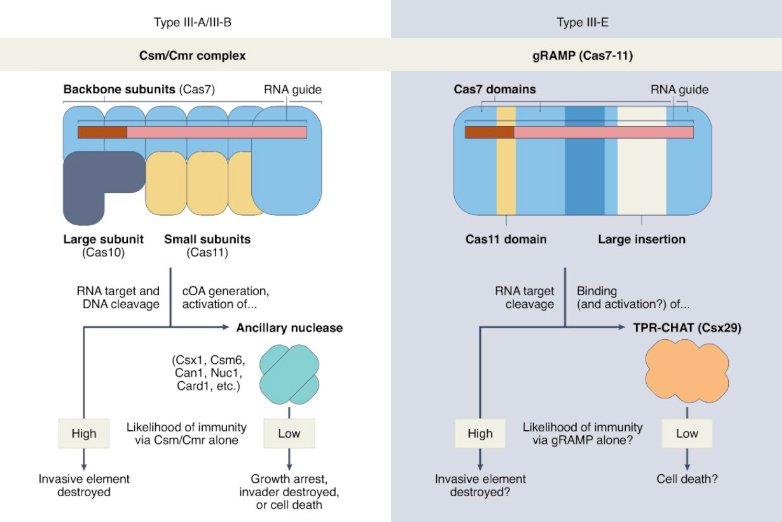
CRISPR meets caspase [News & Views]
Hochstrasser ML & Nuñez JK. Nature Microbiology (2021)
DNA targeting by a minimal CRISPR RNA-guided Cascade
Hochstrasser ML*, Taylor DW*, Kornfeld JE*, Nogales E, Doudna JA. Molecular Cell (2016)
Cutting it close: CRISPR-associated endoribonuclease structure and function [Review]
Hochstrasser ML & Doudna JA. Trends in Biochemical Sciences (2015)
CasA mediates Cas3-catalyzed target degradation during CRISPR RNA-guided interference
Hochstrasser ML*, Taylor DW*, Bhat P, Guegler C, Sternberg SH, Nogales E, & Doudna JA. PNAS (2014)
Genome Editing
Clinical applications of CRISPR‐based genome editing and diagnostics [Review]
Foss DV, Hochstrasser ML, & Wilson RC. Transfusion (2019)
Enhanced genome editing with Cas9 ribonucleoprotein in diverse cells and organisms [Protocol]
Farboud B*, Jarvis E*, Roth TL*, Shin J*, Corn JE, Marson A, Meyer BJ, Patel NH, & Hochstrasser ML. Journal of Visualized Experiments (2018)
COVID-19 Testing
When the U.S. shut down in March 2020 due to the COVID-19 pandemic, the IGI quickly built a pop-up SARS-CoV-2 clinical diagnostic lab and began offering tests to underserved and at-risk Bay Area community members. We later developed a saliva test for asymptomatic UC Berkeley students and staff. As a whole, these efforts helped curb the spread of infection in our area during a time when tests with a quick turnaround were almost entirely unavailable. We released these papers to help other scientists looking to set up testing efforts.
Launching a saliva-based SARS-CoV-2 surveillance testing program on a university campus
Ehrenberg AJ, Moehle EA, Brook CE, Doudna Cate AH, Witkowsky LB, Sachdeva R, Hirsh A, Barry K, Hamilton JR, Lin-Shiao E, McDevitt S, Valentin-Alvarado L, Letourneau KN, Hunter L, Keller A, Pestal K, Frankino PA, Murley A, Nandakumar D, Stahl EC, Tsuchida CA, Gildea HK, Murdock AG, Hochstrasser ML, O'Brien E, Ciling A, Tsitsiklis A, Worden K, Dugast-Darzacq C, Hays SG, Barber CC, McGarrigle R, Lam EK, Ensminger DC, Bardet L, Sherry C, Harte A, Nicolette G, Giannikopoulos P, Hockemeyer D, Petersen M, Urnov FD, Ringeisen BR, Boots M, & Doudna JA. PLoS One (2021)
Blueprint for a pop-up SARS-CoV-2 testing lab
Amen AM, Barry KW, Boyle JM, Brook CE, Choo S, Cornmesser LT, Dilworth DJ, Doudna JA, Ehrenberg AJ, Fedrigo I, Friedline SE, Graham TGW, Green R, Hamilton JR, Hirsh A, Hochstrasser ML, Hockemeyer D, Krishnappa N, Lari A, Li H, Lin-Shiao E, Lu T, Lyons EF, Mark KG, Martell LA, Martins ARO, McDevitt SL, Mitchell PS, Moehle EA, Naca C, Nandakumar D, O’Brien E, Pappas DJ, Pestal K, Quach DL, Rubin BE, Sachdeva R, Stahl EC, Syed AM, Tan I, Tsuchida CA, Tollner A, Tsui CK, Turkalo TK, Urnov FD, Warf MB, Whitney ON, & Witkowsky LB. Nature Biotechnology (2020)
Earlier Work
Spatial quality control bypasses cell-based limitations on proteostasis to promote prion curing ———————————————
Klaips CL, Hochstrasser ML, Langlois CR, & Serio TR. eLife (2014)
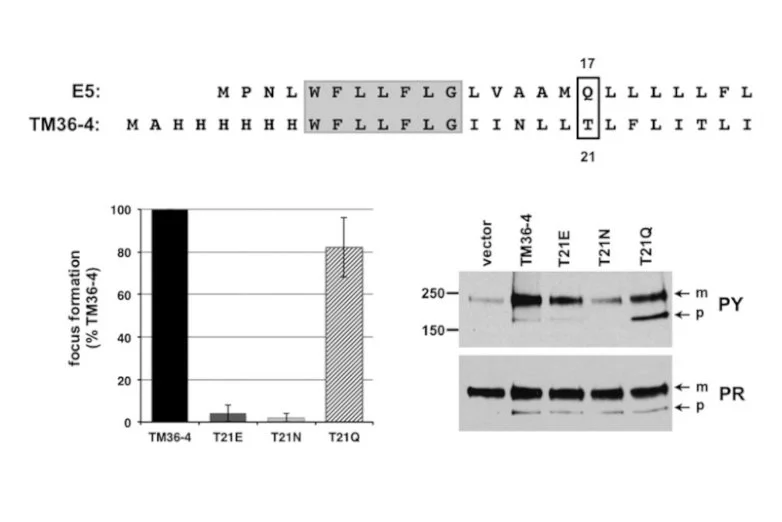
A single amino acid substitution converts a transmembrane protein activator of the platelet-derived growth factor β receptor into an inhibitor
Petti LM, Talbert-Slagle K, Hochstrasser ML, & DiMaio D. JBC (2013)